LAUREN KUEHNE AND JULIAN OLDEN, FRESHWATER ECOLOGY AND CONSERVATION LAB, UNIVERSITY OF WASHINGTON

Figure 1. Mesocosm study where tanks with Brazilian elodea (Egeria densa) and Eurasian milfoil (Myriophyllum. spicatum) were sampled as plants grew and senesced over a 10-week period.
The hope and promise of using environmental DNA (eDNA) – where water samples are analyzed for presence of a species’ DNA – is alluring, and with good reason. Finding the proverbial needle in a haystack is infinitely easier than detecting a fish, invertebrate, or aquatic plant in a lake or river. Unless it’s very abundant, chances are you will miss it! Consequently, tools that can simplify this process, reduce the cost of surveying, or just improve chances of detection are greatly needed. The stakes are even higher in the context of monitoring for invasive species, which require considerable effort and money to manage and control. Aside from prevention, early detection is the best defense, making eDNA a promising and potentially useful tool. In several studies, eDNA has proved equally or sometimes more sensitive than traditional sampling methods, elevating its use to monitor species of high concern such as Asian carp and zebra mussels.
In 2017, we embarked on a research project to assess whether eDNA could be used to detect invasive aquatic plants in lakes. Given that so few studies had been done for plants, we had to start with basic questions:
“Did it matter if plants were actively growing, or if they were dying off?”
“Did detection using eDNA differ between plant species?”
“How important is it to be close to a plant patch to get a detection?”
The million-dollar question, of course, was “How abundant did plants need to be for eDNA detection to work?” – if the answer was “very,” then using eDNA for early detection and rapid response would be problematic.
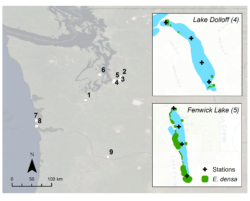
Figure 2. Study lakes in Western Washington with E. densa and/or M. spicatum. Examples of lakes with low and high plant abundance based on visual surveys are shown in insets. From Kuehne et al. 2020 (1)
Supported by a Washington State Department of Ecology Aquatic Invasive Plants Management grant, we implemented the study using both laboratory and field surveys. This combined approach offered the precision and replication afforded by lab conditions to help identify processes, combined with field sampling to allow for a critical real-world reality check. To evaluate how growth stage and species impacted detection using eDNA, we grew two aquatic invasive plants (Brazilian elodea and Eurasian milfoil) in mesocosms over 10 weeks (Figure 1). We collected two one-liter samples from each tank in each week, and partners at the U.S. Geological Survey Western Fisheries Research Center ran assays that they developed for each species. Next, we identified nine lakes in Western Washington that had one or both species present, and varied in plant abundance from those with just a few plant patches up to heavily infested, where plants were present along most of the shoreline (Figure 2). In August, when plant biomass was at its peak, we collected three one-liter samples at four stations in each lake, while mapping presence and abundance of the plants during a visual lake survey. Three of the lakes were also sampled in June and October to assess the importance of sampling timing to detection. Finally, in three lakes, we sampled discrete plant patches, starting at the center and moving outward; this was intended to help answer how important proximity was to detection.
Our results were surprising in many ways. First, although we were able to detect both species using environmental DNA in both laboratory and lake sampling, detection was low compared to what might be expected in a study with animals. Furthermore, as plants grew over the first half of the lab study, detection did not improve reliably, even though there was more biomass in the tanks. The factor that did improve detection was when we turned the lights off after week 5, and plants started senescing. Concentration of eDNA measured in samples was consistently lower for Eurasian milfoil than for Brazilian elodea.
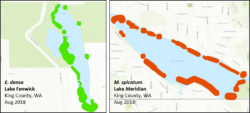
Figure 3. Examples of mapped abundance of Brazilian elodea (green) and Eurasian milfoil (red) in two high abundance lakes where eDNA was reliably detected based on sampling at four stations.
Our field sampling results were highly consistent with the lab results. Based on sampling at four stations, detection using eDNA was reliable only at the highest levels of plant abundance, when plants were effectively found along a majority of the lake shoreline (Figure 3). As in the lab, eDNA concentrations of Brazilian elodea increased by an order of magnitude during fall, when plants were beginning to senesce. And, as in the lab portion of the study, concentrations and detection of milfoil were typically lower than Brazilian elodea, indicating that species may vary considerably in how well eDNA will detect them. Finally, our distance-from-patch sampling showed that being close to patches (and thus the source of eDNA) is important, particularly when overall plant abundance in the lake is low. For example, in a lake with relatively low Eurasian milfoil abundance, the only positive detection was at the edge of the plant patch, even when samples from the center of patches were zero.
Overall, our study showed that much work remains before environmental DNA can be used to detect aquatic invasive plants in lakes. Two of the primary reasons for using eDNA– early detection and evaluation of plant abundance (e.g., following eradication efforts)–are confounded by the problem that detection did not correspond well with plant abundance and was largely unsuccessful unless plants were very abundant. Detection did improve in the fall, indicating that future research might focus on senescence periods. It’s also very important to remember that fewer than ten aquatic plant eDNA studies have been done to date, meaning the application of this technique for plants is still in its infancy. We believe this study offers a cautionary tale in considering eDNA as a “silver bullet” for early detection of plants, but is also an important base from which to systematically examine ways to improve detection in the future.
All results, assays, and data from this study are available online (see sources 1-3 below). You can also email the author for more information related to this work at lauren.kuehne@gmail.com
1. Open Access article of main study and results: Kuehne, L. M., Ostberg, C. O., Chase, D. M., Duda, J. J., & Olden, J. D. (2020). Use of environmental DNA to detect the invasive aquatic plants Myriophyllum spicatum and Egeria densa in lakes. Freshwater Science, 39(3), 000-000. https://www.journals.uchicago.edu/doi/full/10.1086/710106
2. Assay for Brazilian elodea: Chase, D.M., L.M. Kuehne, J.D. Olden, and C.O. Ostberg. 2020. Development of a quantitative PCR assay for detecting Egeria densa in environmental DNA samples. Conservation Genet. Resour. (2020) DOI: https://doi.org/10.1007/s12686-020-01152-w.
3. Online depository of all laboratory and field sampling data. Ostberg, C.O., Kuehne, L.M., Chase, D.M., Duda, J.J., and Olden,J.D., 2020, Detection of invasive aquatic plants Myriophyllum spicatum and Egeria densa in lakes using eDNA, field and mesocosm data: U.S. Geological Survey data release, https://doi.org/10.5066/P90BVKTO
https://www.sciencebase.gov/catalog/item/5e386582e4b0a79317df566a









